A Case for qPCR + NGS
Fungal infection in mastoid cavity
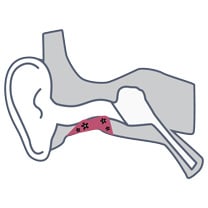
A 79-year-old male with a longstanding history of chronic ear disease has had multiple middle ear and mastoid surgeries, leaving him with a mastoid cavity and non-intact tympanic membrane. He developed an intermittent mastoid cavity infection typically treated with topical ciprofloxacin drops. He then presented with recurrent otorrhea. A sample of the mastoid cavity discharge was collected for MicroGenDX qPCR+NGS testing. This revealed the presence of Aspergillus tamarii — this result was available four days after collection of the specimen. No bacterial DNA was identified, confirming the fungal etiology of the infection. The ciprofloxacin drops were immediately discontinued and topical antifungals used instead with resolution of the infection.
This case demonstrates an important advantages of next-generation DNA sequencing over traditional culture technique, namely time-to-diagnosis. The slow growth of fungi in culture may require more than one week of incubation depending on the species.
Chronic sinusitis requiring dual antibiotic therapy
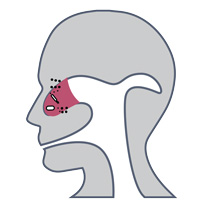
A 57-year-old male with longstanding chronic sinusitis and history of prior endoscopic sinus surgery presented with an exacerbation of his chronic sinusitis. A sinus aspirate was obtained and MicroGenDX next-generation DNA sequencing performed. This revealed medium quantities of bacterial DNA. Three facultative anaerobic organisms were identified: Enterobacter cloacae, Corynebacterium segmentosum, and Staphylococcus epidermidis. The Enterobacter species was found to represent 61% of the bacterial DNA, and the Corynebacterium represented 34%, while S. epi represented only 2% of the DNA – confirming it to be a contaminant. To adequately treat the infection, and given the lack of oral antibiotic coverage with a single agent for both species, dual antibiotic therapy with both doxycycline and ciprofloxacin was used. The patient had complete resolution of infection with one 14-day course of therapy.
This case highlights two important advantages of next-generation DNA sequencing for microbial identification in patients with chronic sinusitis. First, although three bacterial species were identified, the quantitative nature of the results allowed for determination of the relative contribution of each species to the infection, including determining that S. epi was a contaminant. Second, by identifying more than one organism, the need for combined antimicrobial therapy was established, and a treatment with both ciprofloxacin and doxycycline resulted in rapid resolution of infection. The selection of ciprofloxacin and doxycycline would certainly not have occurred as empiric therapy.
Odontogenic sinusitis

A 68-year-old female with a one-year history of sinusitis was previously treated with multiple courses of antibiotics. Her previous ENT obtained an endoscopically directed culture revealing Pseudomonas aeruginosa. Patient was then treated with multiple courses of ciprofloxacin without resolution of her infection. A subsequent culture also revealed scant growth of an anaerobe, Parvimonas micra.
Following additional antibiotics (Augmentin and clindamycin), MicroGenDX next-generation DNA sequencing was performed revealing the presence of six oral anaerobes. A CT of the sinuses was performed that showed total unilateral maxillary sinus opacification and a ruptured periapical abscess of an underlying maxillary molar. Surgery was then performed following extraction of the tooth. Following surgery and additional treatment with clindamycin, her infection resolved.
This case highlights the superior ability of next-generation sequencing to detect the presence of anaerobes compared with culture technique. This allows for the identification of oral anaerobes suggestive of odontogenic sinusitis. Detection of these organisms can then prompt the performance of a CT of the sinuses to detect the odontogenic source. Rapid identification of odontogenic sinusitis can then result in appropriate dental care to treat the source of the infection as well as help direct appropriate antimicrobial therapy.
