MicroGenDX Leads Industry in Published NGS Research
MicroGenDX leads the molecular diagnostics industry in the number of published clinical studies supporting the clinical efficacy of Next-Generation Sequencing diagnostics. To date there have been over 100 published studies across multiple medical specialties utilizing MicroGenDX NGS, with additional studies currently underway. No other molecular laboratory in the U.S. has achieved the same depth and breadth of clinical studies supporting the diagnostic effectiveness of their technology.

An Enhanced Understanding of Culture-Negative Periprosthetic Joint Infection with Next-Generation Sequencing A Multicenter Study
THE JOURNAL OF BONE AND JOINT SURGERY, INCORPORATED 2022 Sep 7;104(17):1523-1529
Study: NGS provides a more comprehensive picture of the microbial profile of infection that is often missed by traditional culture. Examining the profile of PJI in a multicenter cohort using NGS, this study demonstrated that approximately two-thirds of culture-negative PJIs had identifiable opportunistically pathogenic organisms, and furthermore, the majority of infections were polymicrobial. Read Study

Synovial fluid α defensin has comparable accuracy to synovial fluid white blood cell count and polymorphonuclear percentage for periprosthetic joint infection diagnosis
The Bone & Joint Journal Vol. 103-B, No. 6
Article:The aim of this study was to determine the diagnostic accuracy of α defensin (AD) lateral flow assay (LFA) and enzyme-linked immunosorbent assay (ELISA) tests for periprosthetic joint infection (PJI) in comparison to conventional synovial white blood cell (WBC) count and polymorphonuclear neutrophil percentage (PMN%) analysis. Read Article

Reinfection or Persistence of Periprosthetic Join Infection? Next Generation Sequencing Reveals New Findings
Orthopaedic Proceedings
Research Article: Surgical management of PJI remains challenging with patients failing treatment despite the best efforts. An important question is whether these later failures reflect reinfection or the persistence of infection. Proponents of reinfection believe hosts are vulnerable to developing infection and new organisms emerge. Read Article

An Enhanced Understanding of Culture-Negative Periprosthetic Joint Infection With Next-Generation Sequencing
Orthopaedic Proceedings
Research Article: In this prospective study samples were collected in 238 consecutive patients undergoing revision total hip and knee arthroplasties. Of these 83 patients (34.9%) had PJI, as determined using the Musculoskeletal Infection Society (MSIS) criteria, and of these 20 were culture-negative (CN-PJI). Synovial fluid, deep tissue and swabs were obtained at the time of surgery and sent for NGS and culture/MALDI-TOF. Read Article

Cutibacterium acnes is isolated from Air Swabs - Time to Doubt the Value of Traditional Cultures in Shoulder Surgery?
Archives of Bone and Joint Surgery
Research Article: Given high rates of positive Cutibacterium acnes (C. acnes) cultures in cases of both primary and revision shoulder surgery, the ramifications of positive C. acnes cultures remain uncertain. Next generation sequencing (NGS) is a molecular tool that sequences the whole bacterial genome and is capable of identifying pathogens and the relative percent abundance in which they appear within a sample. Read Article

Next-Generation Sequencing Supports Targeted Antibiotic Treatment for Culture Negative Orthopedic Infections
Infectious Disease Society of America 08 September 2022
Article: Bringing molecular diagnostics into practice can provide critical information about the nature of the infective organisms and allow targeted therapy in these otherwise challenging situations. This review article describes the current state of knowledge related to the use and potential of NGS to diagnose infections, particularly in the setting of PJIs. Read Study

The 2018 Definition of Periprosthetic Hip and Knee Infection: An Evidence-Based and Validated Criteria
The Journal of arthroplasty
Research Article: This multi-institutional study of patients undergoing revision total joint arthroplasty was conducted at 3 academic centers. For the development of the new diagnostic criteria, PJI and aseptic patient cohorts were stringently defined: PJI cases were defined using only major criteria from the MSIS definition (n = 684) and aseptic cases underwent one-stage revision for a noninfective indication and did not fail within 2 years (n = 820). Read Article

Mark Coventry Award Human Knee Has a Distinct Microbiome Implications for Periprosthetic Joint Infection
The Journal of Arthroplasty 38 (2023) S2-S6
Study: This prospective study aimed to investigate the presence of a possible microbiome in the human knee and compare the microbial profiles in different knee conditions. Synovial fluid samples were collected from 65 knees with various conditions, including normal knees, osteoarthritis, aseptic revision, and prosthetic joint infection (PJI). The results revealed that there is a distinct knee microbiome, with different microbial compositions observed in different knee conditions. These findings suggest that the knee microbiome may have an impact on the development of osteoarthritis and PJI. Read Study

Tarabichi M, Shohat N, Goswami K, et al. Diagnosis of Periprosthetic Joint Infection: The Potential of Next-Generation Sequencing.
J Bone Joint Surg Am. 2018;100(2):147-154.
Study: A prospective study evaluated the accuracy of next-generation sequencing (NGS) in identifying synovial and deep tissue infective microbes in periprosthetic joint infection patients (65 revision arthroplasties and 17 primary arthroplasties). In infected tissues, culture identified 17 cases and NGS 25 cases. NGS identified microbes in 9 of the aseptic revisions with negative cultures and in 6 of the primary total joint arthroplasties. NGS identified additional organisms that escaped detection via culture methods. Read Study

Semen Microbiota Are Dramatically Altered in Men With Abnormal Sperm Parameters
Nature.com Scientific Reports volume 14, Article number: 1068 (2024)
Study: Unveiling the Microbial Orchestra: Exploring the Intriguing Links Between Semen Microbiome and Sperm Parameters, Offering New Insights into Male Fertility Challenges. Ready Study Here

The Potential of High-Throughput DNA Sequencing of the Paranasal Sinus Microbiome in Diagnosing Odontogenic Sinusitis
Otolaryngol Head Neck Surg. 2019;161(6):1043-1047.
Study: A case series with chart review of 142 patients with sinusitis was performed within a single tertiary care academic medical center. The study objective was to evaluate the use of high-throughput DNA sequencing to diagnose sinusitis of odontogenic origin. The patient’s computed tomography sinus scan was used as a reference standard. In those patients having an odontogenic source based on computed tomography, high-throughput sequencing produced sensitivity and specificity values of greater than 80% in the detection of oral flora species in the sinus. This study supports the use of high-throughput DNA sequencing in supplementing other methods of investigation for identifying an odontogenic etiology of sinusitis. Read Study

Characterization of native knee microorganisms using nextgeneration sequencing in patients undergoing primary total knee arthroplasty
Science Direct
Research Article: A proportion of patients with osteoarthritis undergoing primary total knee arthroplasty have organisms identified in their joint by NGS at the time of surgery. Organisms identified after TKA by NGS when concern for periprosthetic joint infection exists may represent the native microbiome rather than pathogenic microbes. Read Article

The Natural History and Composition of Urinary Catheter Biofilms: Early Uropathogen Colonization with Intraluminal and Distal Predominance
The Journal of Urology 2020 Feb;203(2):357-364
Study: We sought to determine the composition and initiation site of bacterial biofilm on indwelling urinary catheters and to track biofilm progression with time. Indwelling urinary catheters were collected from 2 tertiary care centers following removal from patients. Indwelling time was noted and catheters were de-identified. Catheters were sectioned, stained for biofilms and analyzed by spectrophotometry and visualization. Biofilm colonization patterns were analyzed using descriptive statistical analysis and bacterial composition was determined using next generation sequencing. Read Study

The Role of Bacterial Biofilms and the Pathophysiology of Chronic Rhinosinusitis
Current allergy and asthma reports
Research Article: The earliest description of a bacterial biofilm is likely centuries old. However, only in the past few decades has a wealth of knowledge developed pertaining to this bacterial form of existence. Biofilms have been implicated mainly in chronic disease states, and the current available treatment modalities for infection have demonstrated limited efficacy against bacteria in this form. Read Article

Next-Generation Sequencing Quickly Identifies Mycobacterium smegmatis in Spine Implant Infection - A Case Report
Allergy & Rhinology
Case Report: We present the case of a 62-year-old woman who underwent revision surgery because of screw displacement after C1 to C3 laminectomy and occiput to C5 posterior fusion. Culture of tissue obtained intraoperatively grew nontuberculous mycobacteria (NTM) in 5 days; however, a specific species was not identified using culture until 5 weeks after sample collection. Read Report

Detecting the presence of bacteria in low-volume preoperative aspirated synovial fluid by metagenomic next-generation sequencing
International journal of infectious diseases
Research Article: With a low volume of synovia (1 ml), mNGS can be performed with higher sensitivity and specificity compared to synovial culture. Thus, mNGS can be a useful supplemental method to improve diagnostic efficiency during the preoperative period. Read Article

Fracture-Associated Microbiome and Persistent Nonunion - Next-Generation Sequencing Reveals New Findings
Journal of orthopaedic trauma
Research Article: Positive NGS detection was significantly correlated with persistent nonunion, positive in 77% more cases than traditional culture. Nonunion cases were observed to have significantly increased diversity and altered bacterial profiles from control cases. Read Article

Temporal dynamics of relative abundances and bacterial succession in chronic wound communities
Wound Repair and Regeneration
Research Article: Polymicrobial bacterial infection is an important factor contributing to wound chronicity. Consequently, clinicians frequently adopt a biofilm-based wound care approach, in which wounds are treated utilizing DNA sequencing information about microbial communities. Read Article

Consensus guidelines for the identification and treatment of biofilms in chronic nonhealing wounds
Wound Repair and Regeneration
Research Article: This consensus document attempts to clarify misunderstandings about the role of biofilms in clinical practice, and provides a basis for clinicians to recognize biofilms in chronic nonhealing wounds and manage patients optimally. A new paradigm for wound care, based on a stepped-down treatment approach, was derived from the consensus statements. Read Article

Next-Generation Sequencing for Pathogen Identification in Infected Foot Ulcers
American Orthopaedic Foot & Ankle Society
Research Article: Accurate identification of primary pathogens in foot infections remains challenging due to the diverse microbiome. Conventional culture may show false-positive or false-negative growth, leading to ineffective postoperative antibiotic treatment. Next-generation sequencing (NGS) has been explored as an alternative to standard culture in orthopedic infections. Read Article

Analysis of the chronic wound microbiota of 2,963 patients by 16S rDNA pyrosequencing.
Wound Repair Regen. 2016 Jan-Feb;24(1):163-174.
Research: The study utilized 16S rDNA pyrosequencing to analyze the composition of the bacterial communities present in samples obtained from patients with chronic diabetic foot ulcers (N = 910), venous leg ulcers (N = 916), decubitus ulcers (N = 767), and nonhealing surgical wounds (N = 370). All wound types examined in the study exhibited similar levels of microbial diversity and abundance at a genus level. Neither patient demographics nor wound type influenced the bacterial composition of the chronic wound microbiome. Staphylococcus and Pseudomonas were the two most frequent bacterial genera identified, however significant variability was observed in their abundance. Staphylococcus was the most frequent bacterial genus present in the polymicrobial communities of the chronic wound samples tested; of these, 40% were methicillin‐resistant. Pseudomonas spp. were present in 25% of all wound samples analyzed. Strict anaerobes comprised four of the top 10 genera detected in the chronic wound samples. Read Research

Molecular Microbiology: New Dimensions for Cutaneous Biology and Wound Healing
The Journal of investigative dermatology
Research Article: The role of bacteria in the pathogenesis of chronic, nonhealing wounds is unclear. All wounds are colonized with bacteria, but differentiating colonizers from invading organisms is difficult, if not impossible, at the present time. Furthermore, robust new molecular genomic techniques have shown that only 1% of bacteria can be grown in culture; anaerobes are especially difficult to identify using standard culture methods. Read Article

Torchia MT, Austin DC, Kunkel ST, Dwyer KW, Moschetti WE. Next-Generation Sequencing vs Culture-Based Methods for Diagnosing Periprosthetic Joint Infection After Total Knee Arthroplasty: A Cost-Effectiveness Analysis.
J Arthroplasty. 2019;34(7):1333-1341.
Study: A comparison was made between culture-based techniques and next-generation sequencing (NGS) relating to cost-effectiveness in diagnosing periprosthetic joint infection (PJI) after total knee arthroplasty. NGS is more sensitive than culture-based techniques for identifying microorganisms but is less specific and more expensive. A Markov, state-transition model projecting lifetime costs and quality-adjusted life years (QALYs) was constructed to determine the cost-effectiveness from a societal perspective. The primary outcome was incremental cost-effectiveness ratio, with a willingness-to-pay threshold of $100,000/QALY. Culture was not cost-effective compared to NGS, with an incremental cost-effectiveness ratio of $422,784 per QALY. One-way sensitivity analyses found NGS to be the cost-effective choice above a pretest probability of 45.5% for PJI. The results suggest that NGS should be reserved for clinical contexts with a high pretest probability of PJI. Read Study

Detection of Bacteria by Next-Generation Sequencing in Men with Chronic Prostatitis/Chronic Pelvic Pain Syndrome: Incidence Correlation to Conventional Culture and Impact on Symptoms
The Journal of Urology Vol. 203, No. 4S, Supplement, Sunday, May 17, 2020
Study: Chronic prostatitis/
Chronic pelvic pain syndrome (CPPS) is a syndrome that shares clinical features with urinary infections and a certain subset of patients improve with antibiotics. However, traditional cultures of urine and expressed prostatic secretions often fail to identify an organism. Next-generation sequencing (NGS) analyzes microbial DNA and can identify organisms that fail to grow in traditional cultures. We sought to compare traditional cultures with NGS in men with CPPS and examine the impact on symptoms and treatment response. Read Study

Determining the Utility of Standard Hospital Microbiology Testing: Comparing Standard Microbiology Cultures With DNA Sequence Analysis in Patients With Chronic Sinusitis
World J Otorhinolaryngol Head Neck Surg. 2019;5(2):82-87.
Study: In a prospective cohort study, 50 chronic sinusitis patients underwent standard hospital cultures (SHC) and DNA sequence analysis as a means to identify polymicrobial pathogens. DNA sequence analysis detected 31.9% more microorganisms compared to SHC (P < 0.05). When multiple microorganisms were detected, DNA sequence analysis yielded more positive results compared to SHC (P < 0.05). Culture failed to identify the dominant species in the microbiome 53% of the time; 20% of the patients had anaerobic bacteria detected by DNA analysis missed by SHC. Antimicrobial therapy based on culture results would have been adequate for only 44% of the patients, compared to 74% based on DNA results. In patients with recalcitrant sinus disease, conventional cultures may need to be augmented with or replaced by molecular-based probes. Read Study

Current Recommendations for the Diagnosis of Acute and Chronic PJI for Hip and Knee—Cell Counts, Alpha-Defensin, Leukocyte Esterase, Next-generation Sequencing
Current Reviews in Musculoskeletal Medicine Vol. 11, pages428–438(2018)
Study: Despite significant progress in recent years, the diagnosis of periprosthetic joint infection (PJI) remains a challenge and no gold standard test exists. A combination of serological, synovial, microbiological, histological, and radiological investigations is performed that are expensive, often invasive, and imperfect. Novel biomarkers and molecular methods have shown promise in recent years. The purpose of this review is to provide an update about the diagnostic recommendations for PJI and cover a selection of emerging diagnostic tools Read Study

Molecular diagnostics and personalized medicine in wound care: assessment of outcomes.
J Wound Care. 2011;20(5):232-239.
Retrospective Study: This level A, retrospective cohort study compared healing outcomes in three large cohorts (400-500 patients each) of wound patients managed universally for bioburden: standard of care group, who were prescribed systemic antibiotics on the basis of empiric and traditional culture-based methodologies; treatment group 1, who were prescribed an improved selection of systemic antibiotics based on the results of molecular diagnostics; treatment group 2 who received personalized topical therapeutics (including antibiotics) based on the results of molecular diagnostics. In comparison to the standard of care group, healing rates improved from 48.5% to 62.4% in treatment group 1 and to 90.4% for treatment group 2. The time to complete closure decreased by 26% in treatment group 1 (p<0.001) and 45.9% in treatment group 2 (p<0.001) compared with the standard of care group. Patients in treatment group 2 had >200% better odds of healing at any given time point compared with the other cohorts. Read Study

Metagenomics in diagnosis and improved targeted treatment of UTI
World J Urol. 2020;38:35–43
Review Article: An extensive publication review of metagenomic sequencing’s clinical applications in urological science – including UTI, prophylaxis prior to transrectal prostate biopsy, prostate cancer research, chronic pelvic pain syndrome, urgency urinary incontinence and neurogenic bladder management – and improved targeted treatment results. Read Article

Analysis of Sinonasal Microbiota in Exacerbations of Chronic Rhinosinusitis Subgroups
OTO Open. 2019;3(3):2473974X19875100.
Retrospective Study: In a retrospective review of 134 chronic rhinosinusitis patients, a DNA microbiome analysis was performed in an effort to identify any similarities or differences in clinical subgroups (65 had nasal polyps, 14 had allergic fungal rhinosinusitis, and 55 had neither). Between 1 and 11 taxa were observed in taxa with >2% relative abundance. Samples from nasal polyp patients and allergic fungal rhinosinusitis patients had an increased prevalence of S aureus. Otherwise, bacterial richness was remarkably low in all samples. Read Study

The Microbiome of Diabetic Foot Osteomyelitis
Eur J Clin Microbiol Infect Dis (2016) 35:293–298
Research Article:The purpose of this investigation was to evaluate the diversity of bacteria in diabetic foot osteomyelitis using a 16S rRNA sequencing approach and to compare the results with conventional culture techniques. In this prospective observational study, we obtained 34 bone samples from patients admitted to our hospital with a moderate–severe diabetic foot infection.We analysed the distribution of the 16S rRNA gene sequences in the bone samples, using an Illumina MiSeq Personal Sequencer. We compared the genera that were detected with the cultured pathogens in the bone samples with conventional techniques. Read Article

Can Next Generation Sequencing Play a Role in Detecting Pathogens in Synovial Fluid?
The Bone & Joint Journal 2018 Feb;100-B(2):127-133
Prospective Study:The diagnosis of periprosthetic joint infection can be difficult due to the high rate of culturenegative infections. The aim of this study was to assess the use of next-generation sequencing for detecting organisms in synovial fluid. In this prospective, single-blinded study, 86 anonymized samples of synovial fluid were obtained from patients undergoing aspiration of the hip or knee as part of the investigation of a periprosthetic infection. A panel of synovial fluid tests, including levels of C-reactive protein, human neutrophil elastase, total neutrophil count, alpha-defensin, and culture were performed prior to next-generation sequencing. Read Study

Survey of Fungi and Yeast in Polymicrobial Infections in Chronic Wounds
Journal of Wound Care
Retrospective Study: To assess the incidence, abundance and species diversity of fungi in chronic wounds, as well as to describe the associations of major fungi populations. Comprehensive molecular diagnostic reports were evaluated from a total of 915 chronic wounds in a retrospective study. Read Study
Next Generation Sequencing for Microbial Analysis to Select Prophylactic Antibiotic Selection before Urologic Stone Surgery: A Culture Change
Open Journal of Urology Vol.11 No.7, July 2021
Article: This paper aims to determine if the combination of polymerase chain reaction (PCR) and next-generation sequencing (NGS) could identify bacteria in culture-negative urine that would alter prophylaxis management. Read Article

Reassessment of Routine Midstream Culture in Diagnosis of Urinary Tract Infection
Journal of Clinical Microbiology March 2019 Volume 57 Issue 3
Article: Midstream urine (MSU) culture remains the gold standard diagnostic test for confirming urinary tract infection (UTI). We previously showed that patients with chronic lower urinary tract symptoms (LUTS) below the diagnostic cutoff on MSU culture may still harbor bacterial infection and that their antibiotic treatment was associated with symptom resolution. Read Article

A New Gold Rush: A Review of Current and Developing Diagnostic Tools for Urinary Tract Infections
Diagnostics 2021, 11, 479
Article: Urinary tract infections (UTIs) are one of the most common infections in the United States and consequently are responsible for significant healthcare expenditure. The standard urine culture is the current gold standard for diagnosing urinary tract infections, however there are limitations of the test that directly contribute to increased healthcare costs. Read Article

Next-Generation DNA Sequencing for Infected Genitourinary Implants: How I do it
The Canadian Journal of Urology. 27(5); October 2020
Prospective Study: Infection of artificial urinary sphincters or inflatable penile protheses is one of the most devastating complications after prosthetic surgery and can have a significant impact on a quality of life. Patients undergoing revision surgery with or without device replacement may have increased risk for infection when compared to initial primary surgery. As such, surgeons may utilize traditional culture results to direct antimicrobial therapy for these patients. Unfortunately, culture results can be inconclusive in up to one-third of the time even in the setting of active device infection. Next-generation DNA sequencing (NGS) of DNA is an emerging technology capable of sequencing entire bacterial genomes and has the potential to identify microbial composition in explanted devices. Read Study

Use of Dipstick Assay and Rapid PCR-DNA Analysis of Nasal Secretions for Diagnosis of Bacterial Sinusitis in Children With Chronic Cough
Allergy Rhinol (Providence). 2019;10:2152656718821281.
Study:Twenty-two patients under the age of 15 with chronic cough were assessed with use of nasal dipstick sample acquisition and polymerase chain reaction (PCR)-DNA analysis in an effort to differentiate bacterial sinusitis from other causes of chronic cough and to identify pathogens from the nasal cavity. Potential dominant pathological bacteria were identified in 80% of the children with chronic cough and sinusitis. DNA analysis was an effective tool in identifying relevant pathogens and directing specific therapies. It can be considered as an alternative to imaging studies for sinusitis diagnosis. Read Study

Leptotrichia amnionii, an Emerging Pathogen of the Female Urogenital Tract
Journal of Clinical Microbiology Vol. 45, No. 7
Article: Leptotrichia amnionii, a recently described, very fastidious, gram-negative anaerobic bacterium, is an opportunistic pathogen of the female urogenital tract. We report a case of second-trimester abortion in a patient with chorioamnionitis and L. amnionii bacteremia and a case of renal abscess in a female 5 weeks postpartum. Read Article

Urinary tract and genito-urinary suppurative infections due to anaerobic bacteria
International Journal of Urology 2004 Mar;11(3):133-41
Article: Anaerobes have been involved in many different types of urinary tract infection. This review describes the microbiology, diagnosis and management of urinary tract and genito-urinary suppurative infections caused by anaerobic bacteria. Read Article

The effect of local estrogen therapy on the urinary microbiome composition of postmenopausal women with and without recurrent urinary tract infections
International Urogynecology Journal 2021 May 18
Article: Recurrent urinary tract infections (rUTIs) occur in 2-10% of postmenopausal women. Local estrogen therapy (LET) has been shown to reduce UTIs. This study aimed to compare the urinary microbiome between patients with and without a history of rUTIs and to examine whether treatment with LET influences the diversity and richness of microbiome species in two groups. Read Article

Next generation sequencing in patients with nephrolithiasis: how does it perform compared with standard urine and stone cultures?
Therapeutic Advances in Urology 2021, Vol. 13: 1–8
Article: Our aim was to compare microorganism detection between standard culture (Ctx) and next generation sequencing (NGS) in patients undergoing surgery for nephrolithiasis; we prospectively compared both urine and stone culture results using these two techniques. Read Article

MP77-01 Individual, DNA-Guided, Antibacterial Prophylaxis Prior to Transrectal Prostate Biopsy Based on Results of Next Generation Sequencing (NGS) of Rectal Swabs is a Promising Targeted Approach to Prevent Severe Urinary Tract Infection
Journal of Urology Vol. 201, No. 4S
Article: NGS allows a comprehensive analysis of the genomic profile of rectal microbiota and, most importantly, the identification of resistance genes to the standard antibacterial empiric prophylaxis. The aim of our pilot study was to evaluate NGS of rectal swabs in patients before transrectal biopsy of the prostate, seeking to prevent urinary tract infection (UTI). Read Article

Microorganism Profiles of Penile Prosthesis Removed for Infection, Erosion, and Mechanical Malfunction Based on Next-Generation Sequencing
The Journal of Sexual Medicine 2022 Feb;19(2):356-363
Article: Background: Next-generation sequencing (NGS) is an emerging technology that may allow for more sensitive and sophisticated microbial testing of the microbiota of penile prostheses (PP).
Aim: To describe the microorganism profiles of PP explanted for infection, erosion, and mechanical malfunction using NGS. Read Article

Actinotignum schaalii Infection: A Clandestine Cause of Sterile Pyuria?
Open Forum Infectious Diseases 2018 Feb; 5(2): ofy015
Article: Actinotignum schaalii is an underappreciated cause of urinary tract infections (UTIs) in older adults. The diagnosis may be missed due to difficulty isolating and identifying the organism. Complications can result because the organism is intrinsically resistant to 2 commonly used drugs to treat UTI, as illustrated by this case. Read Article

MP15-14 The value of next generation DNA sequencing testing in rectal swabs before transrectal prostate biopsy for individual and targeted prophylaxis of urinary tract infection
The Journal of Urology. 2018;199(4S):e194. Supplement, Friday, May 18, 2018.
Meeting Abstract: In a phase I study, rectal swabs from 24 patients were collected prior to transrectal prostate biopsy. Based on qPCR + NGS nucleic acid analysis, clinicians provided an individualized, targeted prophylaxis in 15 patients with multidrug resistance (14 patients with fluoroquinolone resistance). In 8 patients, different fungal species were detected, with multi-fungal associations in 4 of these. Subsequent targeted prophylaxis prevented infectious complications in 23 of the 24 patients. Read Abstract

Economic aspects of biofilm-based wound care in diabetic foot ulcers
Journal of Wound Care May 2015, Volume 24, Issue 5
Study: There has been a dramatic rise in the number of chronic wounds globally, which is placing an increased demand on decreasing health-care resources. With significant cuts in health-care budgets, wound care, providers will have to achieve better outcomes quicker and with fewer resources. By using new molecular methods to fully identify wound microbiota, commercially available antimicrobials can be used more efficiently, thereby improving outcomes and decreasing cost. Read Study

Healing and healing rates of chronic wounds in the age of molecular pathogen diagnostics.
J Wound Care. 2010;19(7):272-281.
Study: The study compares healing rates and healing time of two cohorts, one before implementation of molecular diagnostics, one after. Each cohort consisted of ~500 patients with various wound types. The introduction of molecular pathogen diagnostics, with a corresponding precise and targeted therapeutic approach, improved healing rates from 48.5% to 62.4% and reduced healing time by >20% across wound types. Read Study

Goswami K, Parvizi J.
Culture-negative Periprosthetic Joint Infection: Is There a Diagnostic Role for Next-generation Sequencing?. 2020;20(3):269-272.
Culture-negative Periprosthetic Joint Infection: Is There a Diagnostic Role for Next-generation Sequencing?. 2020;20(3):269-272.
Expert Rev Mol Diagn. 2020;20(3):269-272.
Editorial: This review explores the utility of next-generation sequencing (NGS) in diagnosing microbial pathogens in periprosthetic joint infections (PJI). Theoretically, NGS does not suffer from the limitations associated with PCR and culture methods. NGS offers advantages such as processing speed and accuracy, among others. Although several questions regarding the clinical application of NGS remain, review of the literature finds NGS recognizing a greater number of PJI-dwelling microbes and a broader base upon which to apply treatment options. Read Editorial

MP23-06 An utilization of next generation sequencing of urine samples for monitoring of urinary tract infections in patients with neurogenic bladder
The Journal of Urology. 2018;199(4S): e283. Supplement, Friday, May 18, 2018.
Meeting Abstract: In a retrospective study, urine-derived nucleic acid samples were assessed using qPCR + NGS to monitor 13 neurogenic bladder patients. All 13 patients had between 1 and 9 microorganisms present, 10 with a high bacterial load and one with 5 fungal species. Nine patients had antibiotic resistance genes, with 6 having multidrug resistance. Significantly, quinolone resistant E.coli was present in many of the patients, suggesting a required change from empiric therapy to a tailored treatment protocol. Read Abstract

Prophylactic Antibiotics for Prostate Biopsy
AUA News VOLUME 23 | ISSUE 7
Article: The discussion regarding the best strategies for selecting prophylaxis for prostate biopsies to minimize infectious complications continued in this year’s sessions. One group reported on the use of next generation DNA sequencing to test rectal swabs for the purpose of tailoring the prebiopsy antibiotic regimen (MP15-14). Infectious complications were avoided in 23 of 24 patients, with only a single patient having cystitis 3 weeks after biopsy. Read Article

Evaluation of the bacterial diversity of pressure ulcers using bTEFAP pyrosequencing.
BMC Medical Genomics. 2010;3:41-52.
Research Article: Bacterial tag-encoded FLX amplicon pyrosequencing was used to identify the bacterial populations in 49 decubitus ulcers. Decubitus ulcers are shown to be polymicrobial, highly variable (228 genera identified among 49 ulcers) microbial communities dominated by 3 to 10 microorganisms, with obligate anaerobes found in significant proportion. In the decubitus ulcers of diabetics the microbial populations and composition may be significantly different from the communities in non-diabetics. The highly unique profile of each individual wound would require a therapeutic approach tailored to the patient’s respective wound microflora and traditional culture is largely inadequate in determining the microbial composition of many chronic wounds. Read Article

Namdari S, Nicholson T, Abboud J, et al. Comparative Study of Cultures and Next-generation Sequencing in the Diagnosis of Shoulder Prosthetic Joint Infections.
J Shoulder Elbow Surg. 2019;28(1):1-8.
Comparative Study:In a prospective study, routine cultures and next-generation sequencing (NGS) were compared in tissues taken from 44 revision shoulder arthroplasty patients. Defined infection definitions were used when analyzing culture and NGS for concordance. There were no polymicrobial culture results, whereas the NGS results suggested most the infections were polymicrobial. Read Study

An implementation of next generation sequencing for prevention and diagnosis of urinary tract infection in urology
Can J Urol. 2018;25(3):9349-9356.
STUDY: qPCR + NGS was used to identify bacterial and fungal pathogens in 112 UTI patients (acute cystitis, rectal swabs before transrectal prostate biopsy, neurogenic bladder, and chronic bacterial prostatitis). In comparison to culture analysis, qPCR + NGS patients experienced superior targeted treatment outcomes related to detection of resistance genes and anaerobic and culture-independent pathogens. Read Study

Characterizing the Microbiome of the Contracted Breast Capsule Using Next Generation Sequencing
Aesthetic Surgery Journal 2020 Apr 15;sjaa097
Abstract: Recent work suggests that bacterial biofilms play a role in capsular contracture (CC). However, traditional culture techniques provide only a limited understanding of the bacterial communities present within the contracted breast. Next generation sequencing (NGS) represents an evolution of polymerase chain reaction technology that can sequence all DNA present in a given sample. Read Abstract

Evaluation of the bacterial diversity among and within individual venous leg ulcers using bacterial tag-encoded FLX and Titanium amplicon pyrosequencing and metagenomics approaches.
BMC Microbiology. 2009;9:226-237.
Research Article: A pyrosequencing approach was used to characterize the microbial ecology of 40 venous leg ulcer (VLU) samples. Results show that VLU infections are polymicrobial with no single bacterium colonizing the wounds. The most ubiquitous and predominant organisms include previously uncharacterized bacteroidales, various anaerobes, Staphylococcus, Corynebacterium, and Serratia. Topological analysis of VLU show some notable differences in bacterial populations across the surface of the wounds, highlighting the importance of sampling techniques during diagnostics. Metagenomics provide a preliminary indication that there may be protozoa, fungi and possibly an undescribed virus associated with these wounds. Read Article

A Head-to-Head Comparative Phase II Study of Standard Urine Culture and Sensitivity Versus DNA Next-generation Sequencing Testing for Urinary Tract Infections
Rev Urol. 2017;19(4):213–220.
STUDY: 44 acute cystitis patients and 22 control group patients were treated based on either C&S or qPCR + NGS DNA sequencing results. The acute cystitis patients and the control group patients were randomly assigned to be treated either based on culture results or qPCR + NGS DNA sequencing results. On day 14, symptom scores from those treated based on qPCR + NGS DNA sequencing results were statistically better than for those treated based on culture results. Read Study

A Pilot Study to Evaluate Next Generation DNA Sequencing Testing to Rectal Swabs
European Urology Supplements Volume 16, Issue 10, November 2017, Page e2656
Meeting Abstract: The introduction of next generation sequencing (NGS) via DNA technology allows us to analyze the complete genomic profile of the gut microbiota with detection of resistance genes to the most frequently used antibiotics in empiric prophylaxis. The aim of our study was to evaluate NGS of rectal swabs prior to transrectal prostate biopsy to prevent infectious complications. Read Abstract

Survey of bacterial diversity in chronic wounds using Pyrosequencing, DGGE, and full ribosome shotgun sequencing.
BMC Microbiology. 2008;8:43-58.
Research Article: A survey was conducted of major bacterial populations which occur in the pathogenic biofilms of three chronic wound types: diabetic foot ulcers (D), venous leg ulcers (V), and pressure ulcers (P). The study used 3 separate 16S-based molecular amplifications followed by pyrosequencing, shotgun Sanger sequencing, and denaturing gradient gel electrophoresis. The survey study revealed that a wide variety of bacteria with different physiological and phenotypic preferences are common as part of pathogenic biofilm communities in chronic wounds. However, different types of wounds may have different bacterial populations that are prevalent, such as pressure ulcers in which 62% of the populations were identified as obligate anaerobes. Culturing failed to identify major contributing populations, especially strict anaerobes, within the given wound types. Read Article

Corona PS, Goswami K, Kobayashi N, et al. General Assembly, Diagnosis, Pathogen Isolation: Proceedings of International Consensus on Orthopedic Infections.
J Arthroplasty. 2019;34(2S):S207-S214.
Summary: 85% of assembly delegates agree that molecular techniques such as next-generation sequencing (NGS) have been shown to be powerful tools in detecting prosthetic joint infections (PJIs) with negative cultures. Read Summary

Microbiota is a primary cause of pathogenesis of chronic wounds.
J Wound Care. 2016;25(Sup10):S33-S43.
Research Article: 35 out of 43 (81%) wound microbiota harvested from patients produced similar chronic wounds (producing slough and exudate) in a murine chronic wound model. The top 30 species present in patient wounds were identified by molecular sequencing in the seeded mouse wounds. These results demonstrate that wound microbiota actively disseminates from the chronic wound by forces and mechanisms intrinsic to the wound. Koch’s postulates and Hill’s criteria for causation together suggest chronic wound microbiota to be the main cause underlying the pathogenesis of chronic wounds. Read Article

In vitro multispecies Lubbock chronic wound biofilm model.
Wound Repair and Regeneration. 2008;16(6):805-813.
Research: The study addresses the need for a chronic pathogenic biofilm laboratory model that allows for cooperative growth of three commonly seen organisms: Pseudomonas aeruginosa, Enterococcus faecalis, and Staphylococcus aureus. The study developed a novel media formulation, simple laboratory system, quantitative polymerase chain reaction for monitoring population dynamics, and methods for objectively and subjectively measuring biofilm formation. The 24‐hour model reacts as predicted using known biofilm effectors, lending itself to future work in the development and testing of first‐generation chronic wounds pathogenic biofilm therapeutics. Read Research

Goswami K, Tarabichi M, Shohat N, et al. Next Generation Sequencing for the Diagnosis of Periprosthetic Knee Infection: A Multicenter Investigation.
AAHKS 2018 Annual Meeting press release; Dallas, November 3, 2018.
Press Release: A multi-institutional study examined the ability of next-generation sequencing (NGS) to identify the causative organism(s) in patients with PJI of the knee. Recent reports demonstrate that NGS facilitates pathogen identification in culture-negative PJI (CN-PJI); however, this signal has not been externally corroborated. From 13 academic institutions, samples from 102 TKA revisions were examined via NGS. NGS identified microbes in 9 of 10 culture-negative patient samples and 25 of 66 “aseptic” revisions; culture identified microbes in 4 of 66 samples. NGS was able to detect a pathogen in >90% of culture-negative cases and demonstrated a high rate of concordance with culture in culture-positive cases. The collaborative findings suggest NGS is a useful adjunct for identifying the causative organism in PJI, particularly in the setting of CN-PJI. Read Press Release

The presence of biofilm structures in atherosclerotic plaques of arteries from legs amputated as a complication of diabetic foot ulcers.
J Wound Care. 2016;25(2):S16-S22.
Research Article:The study used Fluorescence microscopy and fluorescence in situ hybridization (FISH) to identify potential inflammatory bacterial and viral components of biofilm in atherosclerotic arteries as an investigation into the cause of the chronic inflammation within atherosclerotic plaques of arteries from legs amputated as a complication of diabetic foot ulcers. The results indicate that the presence of biofilms in grossly involved arteries may be an important factor in chronic inflammatory pathways of atherosclerotic progression, in the amputated limbs of patients with diabetic foot ulcers and vascular disease. Read Article

Polymicrobial Nature of Chronic Diabetic Foot Ulcer Biofilm Infections Determined Using Bacterial Tag Encoded FLX Amplicon Pyrosequencing (bTEFAP).
PLoS One. 2008;3(10):e3326.
Research Article: A new bacterial tag encoded FLX amplicon pyrosequencing (bTEFAP) approach was used to investigate the polymicrobial nature of chronic diabetic extremity ulcer infections in 40 patients. The most prevalent bacterial genus associated with diabetic chronic wounds was Corynebacterium spp. Also ubiquitous were obligate anaerobes including Bacteroides, Peptoniphilus, Fingoldia, Anaerococcus, and Peptostreptococcus spp. Other major components of the bacterial communities included commonly cultured genera such as Streptococcus, Serratia, Staphylococcus and Enterococcus spp. We introduce the concept of functional equivalent pathogroups (FEP) as consortia of genotypically distinct bacteria that symbiotically produce a pathogenic community. Together, individual members may synergistically cause disease when in an FEP, giving the biofilm community the factors necessary to maintain chronic biofilm infections. The results suggest that traditional culturing methods may be an extremely biased diagnostic tool, as they select for easily cultured organisms such as Staphylococcus aureus and against difficult to culture bacteria such as anaerobes, which data suggests are ubiquitous in these diabetic extremity ulcer chronic wounds. Read Article

Tarabichi M, Shohat N, Goswami K, Parvizi J. Can Next Generation Sequencing Play a Role in Detecting Pathogens in Synovial Fluid?.
Bone Joint J. 2018;100-B(2):127-133.
Study: In a prospective, single-blinded study, periprosthetic joint infection was assessed in 86 synovial fluid samples from patients undergoing aspiration of the hip or knee. Next-generation sequencing (NGS) identified periprosthetic microbial pathogens in both culture-positive and culture-negative samples of synovial fluid. Unlike culture results, NGS detected antibiotic resistant pathogens in four cases; in 10 culture-negative samples, NGS identified positive results. Read Study

Comparison of culture and molecular identification of bacteria in chronic wounds.
Int J Mol Sci. 2012;13(3):2535-2550.
Comparative Retrospective Study: A total of 168 chronic wounds were studied, using both aerobic bacterial cultures and culture-free DNA analysis. Seventeen (17) different bacterial taxa were identified with culture, and 338 different bacterial taxa were identified with molecular testing. The majority of bacteria identified with the molecular testing were not identified with culture testing; many of these bacteria were obligate anaerobes. In the majority of the samples, culture underreported the diversity of the wound microbiota and failed to detect the most abundant bacteria in the wound. This study demonstrates the increased sensitivity that molecular microbial identification can have over culture methodologies. Read Study

Fungal Nails? DNA Facts Challenge Dystrophic Etiology
Journal of the American Podiatric Medical Association Volume 122: Issue 2
STUDY: Historically recalcitrant to treatment, infection of the nail unit is a pervasive clinical condition affecting approximately 10% to 20% of the US population; patients present with both cosmetic symptomatology and pain, with subsequent dystrophic morphology. To date, the presumptive infectious etiologies include classically reported fungal dermatophytes, nondermatophyte molds, and yeasts. Until now, the prevalence and potential contribution of bacteria to the clinical course of dystrophic nails had been relatively overlooked, if not dismissed. Previously, diagnosis had largely been made by means of clinical presentation, although microscopic examinations (potassium hydroxide) of nail scrapings to identify fungal agents and, more recently, panel-specific polymerase chain reaction assays have been used to elucidate causative infectious agents. Each of these tools suffers from test-specific limitations. Using DNA sequencing-based diagnostics, the results in this article document the first identification and quantification of significant bacterial, rather than mycotic, pathogens to the clinical manifestation of dystrophic nails. In direct opposition to the prevailing and presumptive mycotic-based causes, the results in this article invoke questions about the very basis for our current standards of care, including effective treatment regimens. Read Study

Tarabichi M, Alvand A, Shohat N, Goswami K, Parvizi J. Diagnosis of Streptococcus canis Periprosthetic Joint Infection: the Utility of Next-generation Sequencing.
Arthroplast Today. 2017;4(1):20-23.
Case Report: A 62 year old man presented with an infected prothesis from a primary knee arthroplasty three years prior. Next-generation sequencing identified the infective organism as Streptococcus canis, which cultures failed to detect. The patient was successfully treated and later underwent reimplantation arthroplasty. Read Report

Clinical identification of bacteria in human chronic wound infections: culturing vs. 16S ribosomal DNA sequencing.
BMC Infect Dis. 2012;12:321-329
Comparative Study: Parallel samples from 51 chronic wounds were studied using aerobic culturing and 16S DNA sequencing for the identification of bacteria. Compared to 16S DNA sequencing, aerobic culture detected only half of the bacteria determined to comprise a dominant portion of the microbiota. At the genus level, molecular testing identified 85% (78/92) of the bacteria identified by culture. Conversely, culturing detected 15.7% (78/497) of the aerotolerant bacteria and detected 54.9% of the collective aerotolerant relative abundance of the samples. Read Study

MP71-03 Comparison of Novel DNA-Based Tools for Bacterial Detection in Patients with Chronic Urinary Tract Infection (CUTI)
The Journal of Urology. 2019;201(4S):e1046-e1047. Supplement, Sunday, May 6, 2019.
Meeting Abstract: In a prospective, feasibility, bi-institutional study, four methods of chronic UTI microbiome identification were compared in a population of 74 chronic UTI patients using C&S, extended C&S + PCR, Volente Diagnostics, and MicroGenDX qPCR + NGS. A significant difference in accuracy was observed between C&S versus MicroGenDX. MicroGenDX identified microbes in 17 culture negative patients, and identified causative pathogens in 67 of 69 patients. The MicroGenDX methodology provided the most accurate detection of UTI pathogens over other methods. Read Abstract

Next Generation Sequencing of Microbiomes in Post-DRE Urine Samples of Prostate Cancer Patients
University of CO 2019
ABSTRACT: Pathogenic microorganisms could be responsible for asymptomatic and symptomatic inflammatory processes in the prostate including Escherichia coli, Pseudomonas spp., Neisseria gonorrhoeae, Trichomonas vaginalis, etc (1). Modifications of bacterial populations were observed according to different inflammatory and tumor conditions in prostate cancer (PCa) samples, which may promote the development of cancer by enhancing extracellular environmental factors within the prostate (2-4). Most of these studies have used polymerase chain reaction (PCR) techniques to identify specific microorganisms. Next-generation sequencing (NGS) has much higher sensitivity than PCR to identify microorganisms in biological samples. We investigated microorganisms in post-digital rectal exam (DRE) urine samples obtained from PCa patients and a screening population with a PSA < 1.5 ng/mL using NGS and PCR techniques. Read Abstract
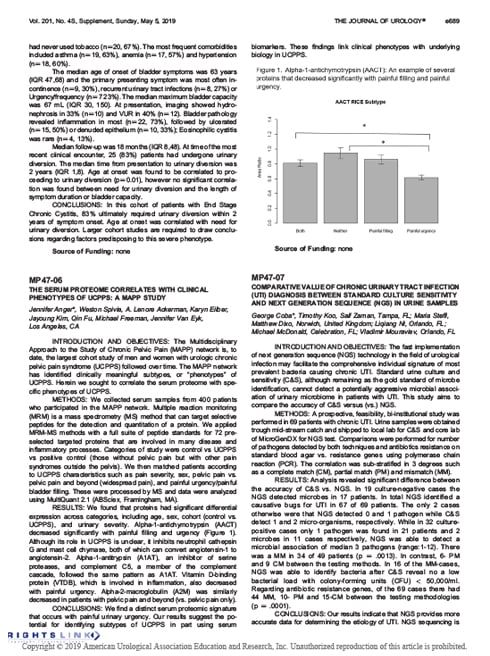
MP47-07 Comparative Value of Chronic Urinary Tract Infection (UTI) Diagnosis Between Standard Culture Sensitivity and Next Generation Sequence (NGS) in Urine Samples
The Journal of Urology. 2019;201(4S):e689. Supplement, Sunday, May 5, 2019.
Meeting Abstract: Using urine samples, the accuracy of C&S versus qPCR + NGS was compared in a 69 UTI patients in a prospective, feasibility, bi-institutional study. C&S failed to identify microbes in 17 infected patients. When compared, NGS test results were more accurate than C&S (97% versus 46%) and revealed a greater presence of polymicrobial pathogens. Read Abstract

Fungal Diversity and Onychomycosis
Arthroplasty Today 2017 Nov 3;4(1):20-23
STUDY: An Analysis of 8,816 Toenail Samples Using Quantitative PCR and Next-Generation Sequencing. Onychomycosis is a fungal infection of the nail that is often recalcitrant to treatment and prone to relapse. Traditional potassium hydroxide and culture diagnosis is costly and time-consuming. Therefore, molecular methods were investigated to demonstrate effectiveness in diagnosis and to quantify the microbial flora present that may be contributing to disease. Read Study

An in vivo polymicrobial biofilm wound infection model to study interspecies interactions.
PLoS One. 2011;6(11):e27317.
Research Article: Multispecies biofilms consisting of both Gram negative and Gram positive strains, as well as aerobes and anaerobes, were grown in vitro and then transplanted onto the wounds of mice. Wounded mice given multispecies biofilm infections displayed a wound healing impairment over mice infected with a single-species of bacteria. In addition, the bacteria in the polymicrobial wound infections displayed increased antimicrobial tolerance in comparison to those in single species infections. These data suggest that synergistic interactions between different bacterial species in wounds may contribute to healing delays and/or antibiotic tolerance. Read Article

Sinus Culture Poorly predicts Resident Microbiota
Int Forum Allergy Rhinol. 2015;5(1):3-9.
Research Study: In a prospective case series, samples from 54 chronic rhinosinusitis patients, collected after surgery, were analyzed by both clinical culture and 16S ribosomal RNA (rRNA) gene sequencing. The accuracy of clinical culture compared to DNA-based molecular techniques was assessed. Each subject had an average of 3 isolates identified by culture and 21.5 ± 12.5 species identified by 16S sequencing. Dominant taxa (>10% abundance) detected via sequencing were detected with culture less than 50% of the time. Only 29.5% of the cultured isolates represented the dominant species. Read Study
