Next-generation sequencing is more effective, efficient, and designed to put patient well-being first compared to microbiology cultures.
In healthcare, efficacy and speed are cardinal factors determining courses of analysis, diagnosis, and treatment. However, infectious disease diagnostics, microbiological culture technology — a method dating back 160 years— is still the standard method for determining types and quantities of organisms. However, next-generation sequencing (NGS) is a newer and more accurate method employed by MicroGenDX to generate accurate life-changing — and often life-saving — results in a short timeframe to better suit patient outcomes.
Originating from advancements in DNA sequencing, MicroGenDX NGS takes a comprehensive approach to DNA sample analysis, with the ability to detect and match more than 50,000 species of bacteria and fungi in its database, the largest of its kind in the world. Paired with a polymerase chain reaction (PCR) panel that amplifies targeted DNA segments, NGS’s thorough analysis capabilities bring centuries-old culture-based diagnostics into the 21st-century by equipping physicians with a technology that emphasizes not just speed and breadth of analysis but also the efficacy of precision treatments and the overall health of patients.
Since its introduction into the healthcare field in the early 2000s, NGS has seen widespread use among physicians specializing in practice and study such as infectious diseases, microbiology, urology, wound care, and ENT.
“Clinicians know that rapid diagnostic technologies are available,” says Dr. James Snyder, director of Microbiology and Infectious Disease Molecular Diagnostics at Louisville Hospital. “They are demanding it, and we are obligated to provide it. If you do not adapt and understand new technologies, you will not survive doing what we do. I love [NGS], and I see what we can do with it…to help our clinicians and not hinder them, and ultimately to promote the appropriate use of antibiotics.”
Microbiology Cultures vs. NGS
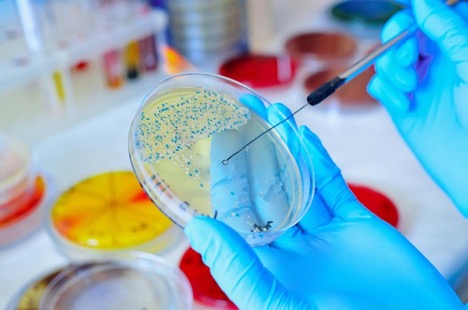 Microbial culture technology is widely used and widely known — students are introduced to it as early as high-school biology class. The practice involves introducing a swabbed sample into a petri dish containing a pre-determined culture medium to promote cell growth. However, despite its wide adoption and continued use, microbiology cultures provide underwhelming results compared to NGS.
Microbial culture technology is widely used and widely known — students are introduced to it as early as high-school biology class. The practice involves introducing a swabbed sample into a petri dish containing a pre-determined culture medium to promote cell growth. However, despite its wide adoption and continued use, microbiology cultures provide underwhelming results compared to NGS.
According to a study in The World Journal of Otorhinolaryngology-Head and Neck Surgery, sequencing detected 31.9% more microorganisms than traditional hospital cultures (SHC). Sequencing also yielded more positive results when multiple organisms were detected.
So why do microbiology cultures fail or underperform when compared to NGS? For issues like chronic infections, the root cause is often polymicrobial. Additionally, the culture that does grow might not represent the dominant organisms in a provided sample.
“Relying on the bacterial culture that often yields only the most dominant organism or only a single one is unlikely to lead to appropriate antimicrobial therapy and the resolution of infection,” notes the study. “Therefore, when the SHC results produced a dominant isolate, they often failed to detect other potential pathogens that may contribute to the infectious microbiome.”
Benefits of MicroGenDX’s Next-Generation Sequencing
- Ability to match microbes detected in samples to a database of 50,000 species
- Least expensive cost worldwide for qPCR + NGS testing
- Accepts the widest variety of sample/media types
- Faster turnaround times compared to microbiology cultures
- qPCR detection of 17 antimicrobial resistance genes for all major classes of antimicrobials
MicroGenDX’s NGS proves the more effective process due to its ability to sequence up to millions of DNA strands, identifying all species of bacteria and fungi. Viruses, parasites, and antibiotic resistance genes are detected using PCR-based assays, making the totality of MicroGenDX’s molecular methods more comprehensive in terms of the scale of detection and analysis.
“The major shortfall of biomarkers is that they diagnose the problem, but they do not tell you what the infecting organism is,” says Dr. Javad Parvizi, director of clinical research at the Rothman Institute. “Next-generation sequencing looks at the quantity of DNA that’s in the samples, will run that against the phylogeny of bacteria and fungi, and then it will tell you which bacteria or fungi exist there.”
MicroGenDX accepts the following samples:
- Synovial fluid
- Hardware/heart valve
- Blood
- Urine
- Cerebrospinal fluid (CSF)
- Sputum and saliva
- Respiratory swabs
- Bronchoalveolar lavage (BAL)
- Periodontal pocket samples
- Tissue biopsy, drainage, swab
- Finger and Toenails
It is Time to Modernize Standard Hospital Testing
 To date, roughly 80,000 medical professionals have relied on MicroGenDX’s NGS services to identify and treat chronic bacterial and fungal infections in patients, and that number is growing. Since its inception, MicroGenDX has completed more than 1.5 million molecular tests, including NGS and qPCR, with a certified CAP proficiency rate of 99.2%. However, as effective as NGS is, the adoption throughout the medical profession has been slow.
To date, roughly 80,000 medical professionals have relied on MicroGenDX’s NGS services to identify and treat chronic bacterial and fungal infections in patients, and that number is growing. Since its inception, MicroGenDX has completed more than 1.5 million molecular tests, including NGS and qPCR, with a certified CAP proficiency rate of 99.2%. However, as effective as NGS is, the adoption throughout the medical profession has been slow.
Hospitals currently employing microbiology cultures face lab inefficiencies which can bog down hospital operations. Diagnostic test results are inherently variable, with turnaround times that range from days to weeks. They are also less effective, with a 50% chance of “no growth” results and a less than 30% chance of determining a dominant species. As a result, treatments are often “best guess” antimicrobial prescriptions that cause repeat office visits for patients.
On the other hand, in conjunction with PCR, NGS have rapid turnaround times that range from three to five days, boasting a 99% accuracy rate at identifying all bacteria and fungi in a sample while detecting potential antimicrobial resistance genes. That level of effectiveness informs research-based precision treatment that ends with patient relief and lower overall costs for treatment.
MicroGenDX is committed to its role in helping clinicians identify and treat chronic infections through next-generation sequencing. When laboratory methods fail to identify fastidious or slow growth organisms, NGS direct identification is the gold standard.
At MicroGenDX, this includes:
- Direct colony identification from agar and broth media (including positive blood cultures) for bacteria, mycobacteria, and fungi
- Direct identification of bacteria, mycobacteria, and fungi from specimens
- Identification of up to 17 clinically significant antimicrobial resistance genes in Gram-positive and Gram-negative organisms
- Widest selection of specimen types
For more information, visit microgendx.com/next-generation-sequencing-process.



